dbSNO 3.0 Tutorial
Table of Contents
Tutorial in Search Page
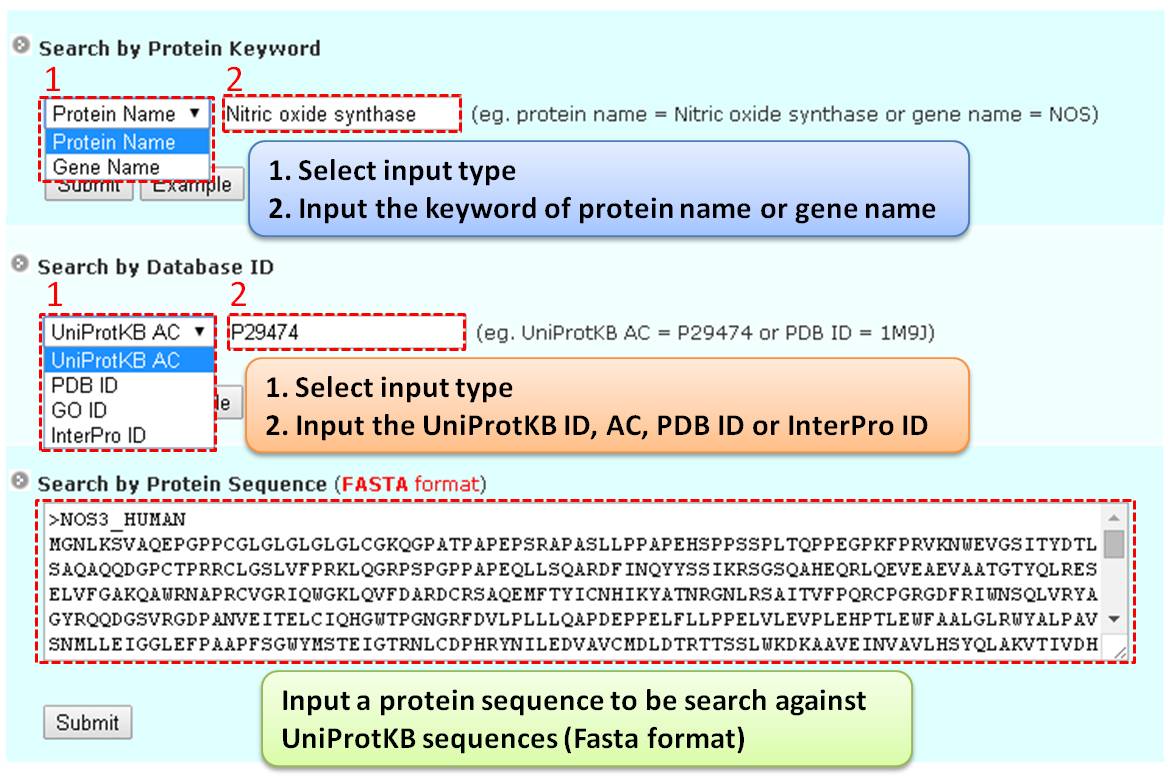
Step 1: Three major queries are available on the search page. Users can query their interested protein by inputting the protein name, gene name, UniProtKB ID, AC, PDB ID, or InterPro ID in the search field of the homepage. It also allows querying amino acid sequences against the release 2025-06 of Swiss-Prot protein sequences.

Step 2: The search results display proteins matching the query keywords in a table. Users can click the "show" URL link to view detailed information about the protein, such as function, S-nitrosylation sites, protein domains, etc.
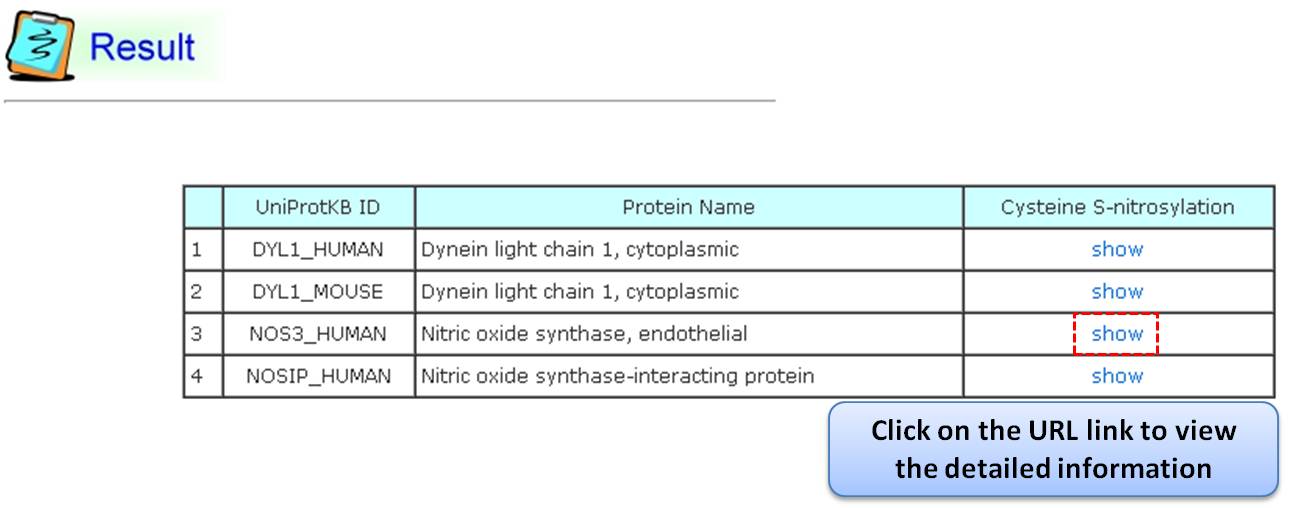
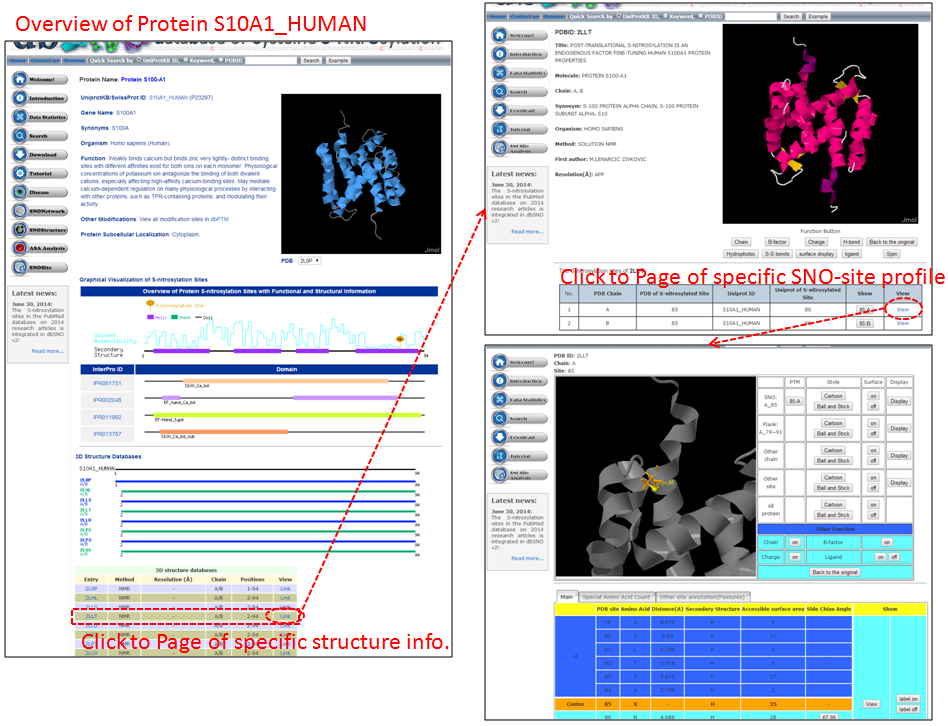
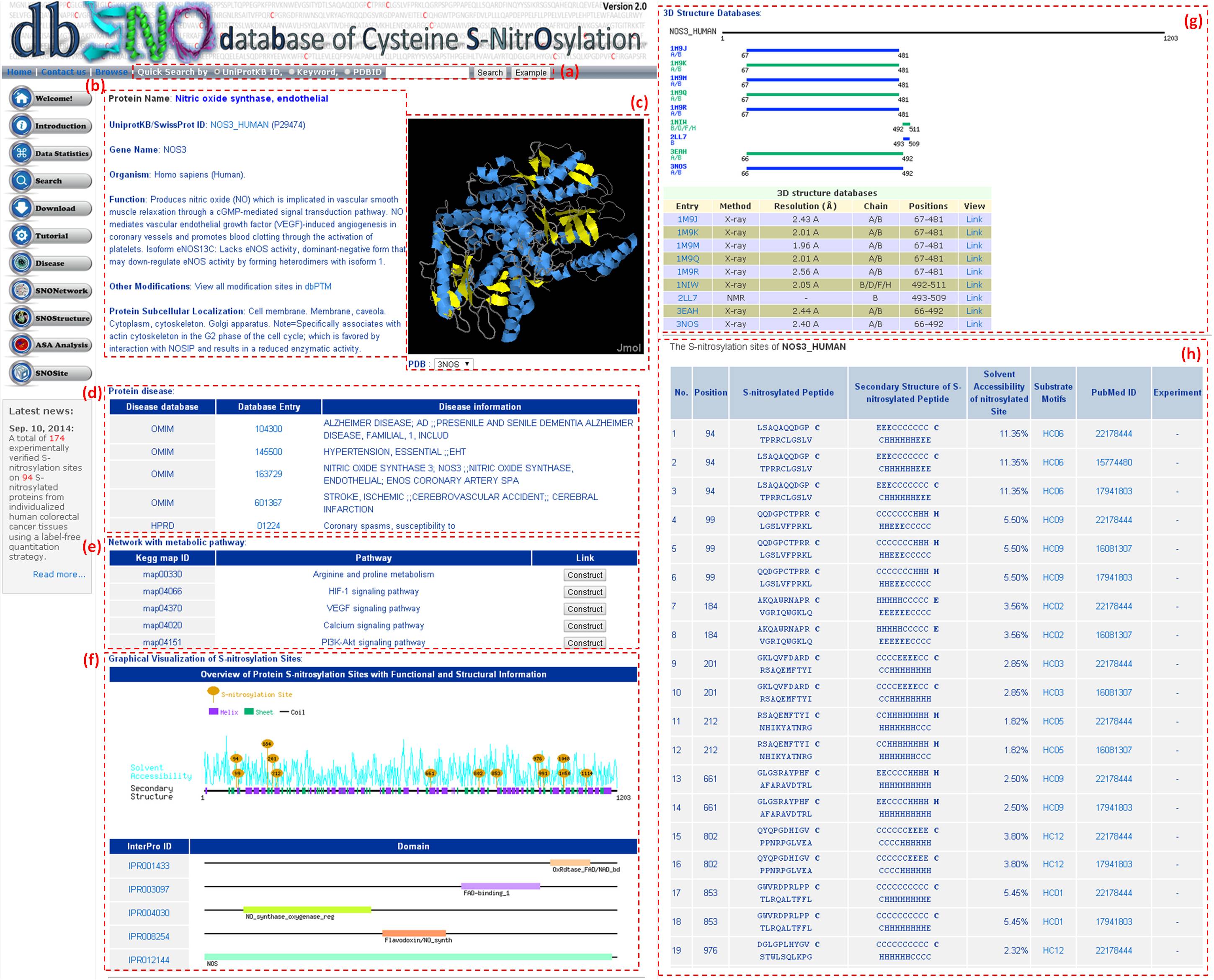
Step 3: Detailed information about S-nitrosylation includes experimental sites and graphical visualizations. Visualizations provide an overview of S-nitrosylation sites, solvent accessibility, protein variations, secondary structures, and functional domains. (a) Quick search by IDs and keywords. (b) Basic information. (c) Visualization of tertiary structure. (d) Protein disease information. (e) Network with metabolic pathway. (f) Graphical visualization of S-nitrosylation sites. (g) NO Site Structure Analysis. (h) Table of S-nitrosylation sites with reported literatures.

Step 4: Users can investigate whether a S-nitrosylation site is located in conserved substrate motifs by clicking the URL in the S-nitrosylation sites table.
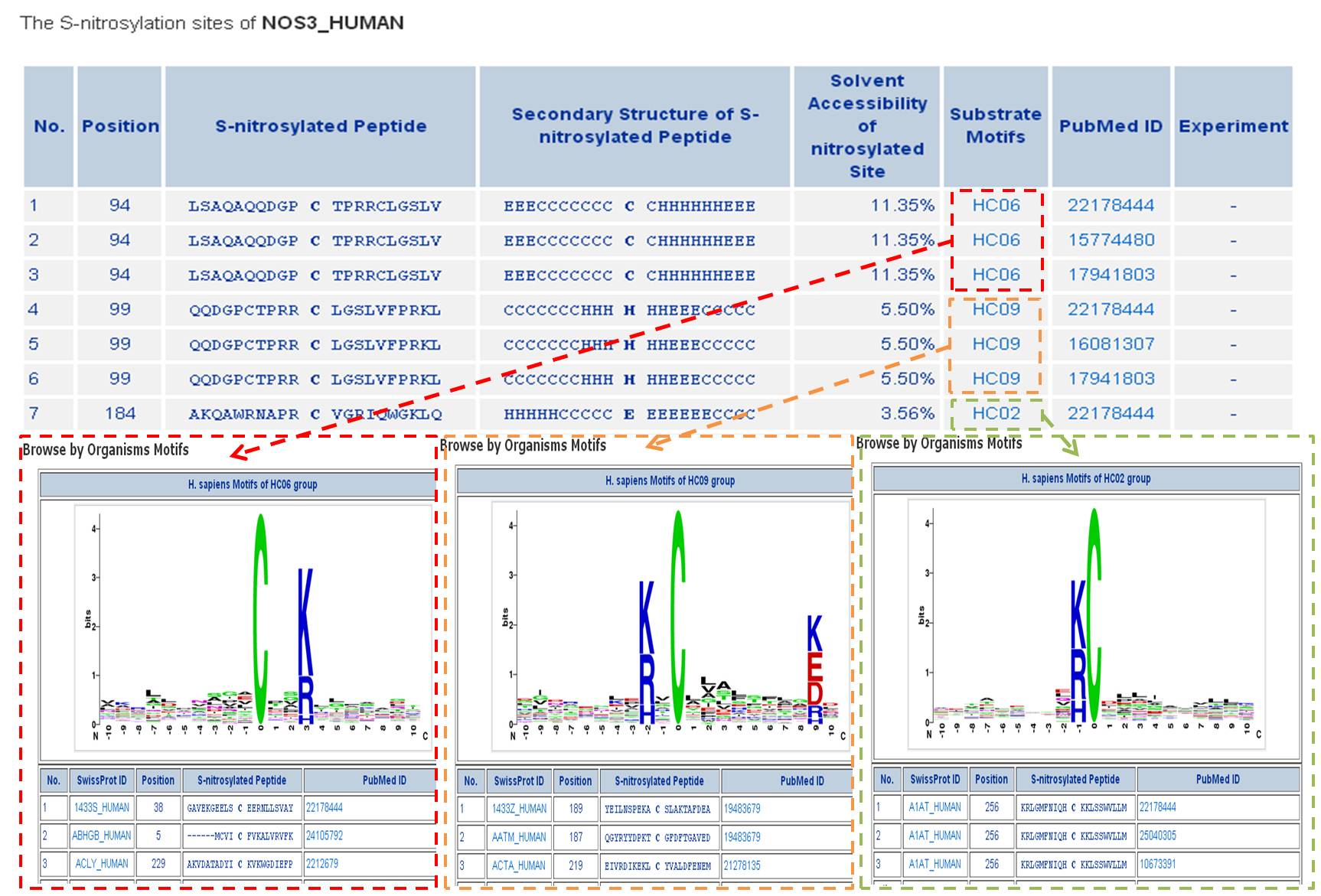
Tutorial in Data Statistics Page
Step 1: The summary table of S-nitrosylation facilitates investigation by category. It lists all S-nitrosylation types in dbSNO 3.0, categorized by modified amino acids and the number of verified sites. For example, users can select M. musculus (Mouse) for detailed information such as modification positions, sequence locations, chemical formulas, and mass differences.
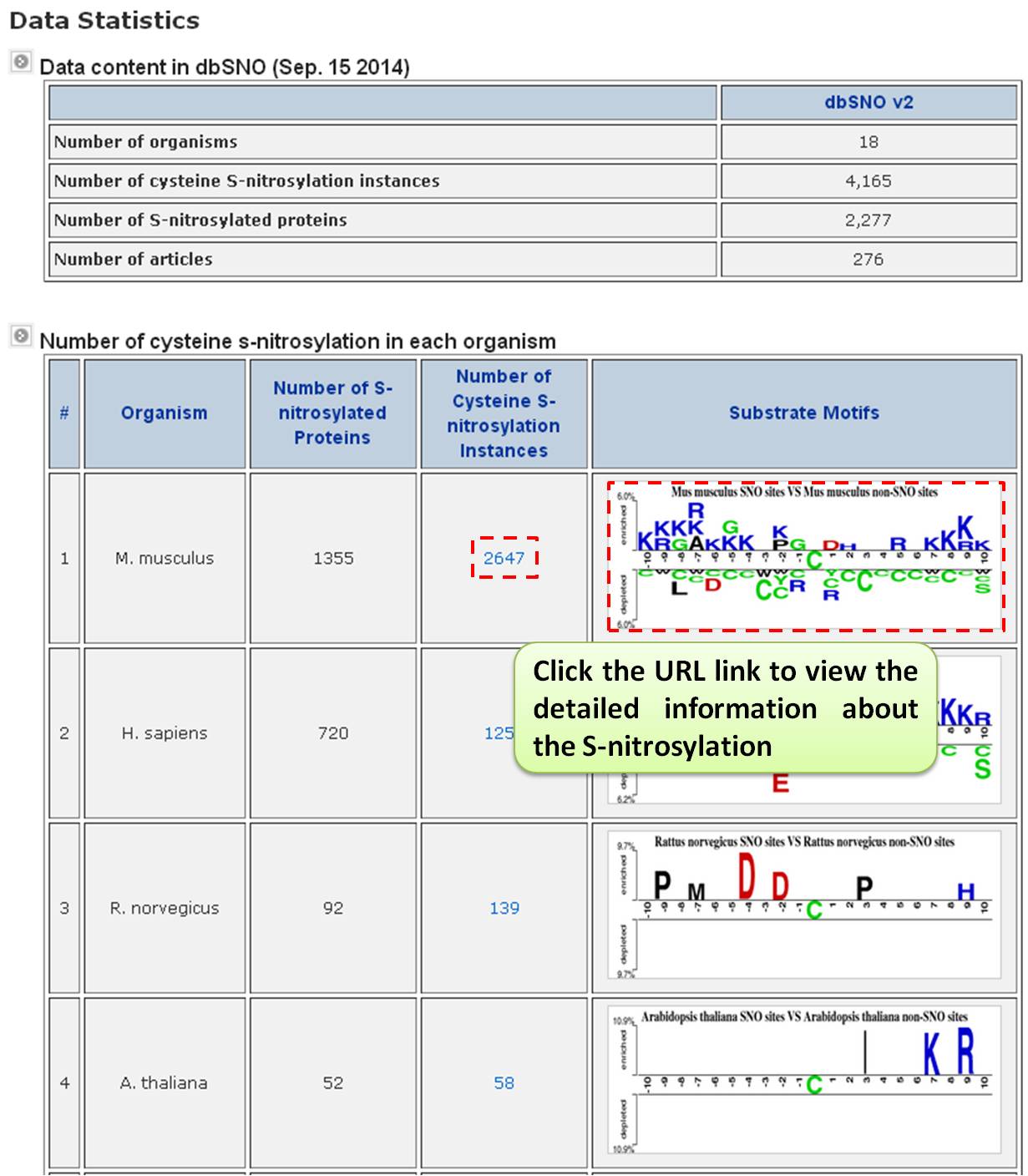
Step 2: Users can explore detailed information for M. musculus (Mouse), including modification positions, sequence locations, chemical formulas, and mass differences. The formula structure is visualized using the Jmol program (JAVA Applet). Key insights include substrate site specificity, such as amino acid frequency, average solvent accessibility, and secondary structure frequency, with sequence logos provided.

Tutorial in SNOStructure Page
This page uses structure alignment methods to compare differences between N=O binding and N=O non-binding in the same protein and SNO site, assessing the effect of N=O binding on protein structure.
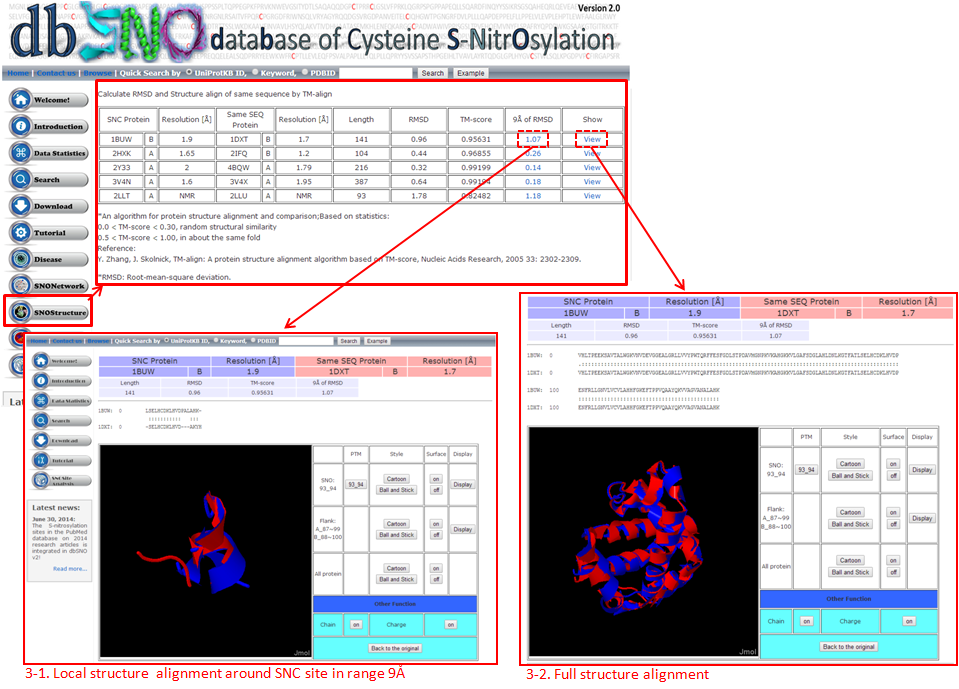
3-1. Local Structure Alignment (9Å)
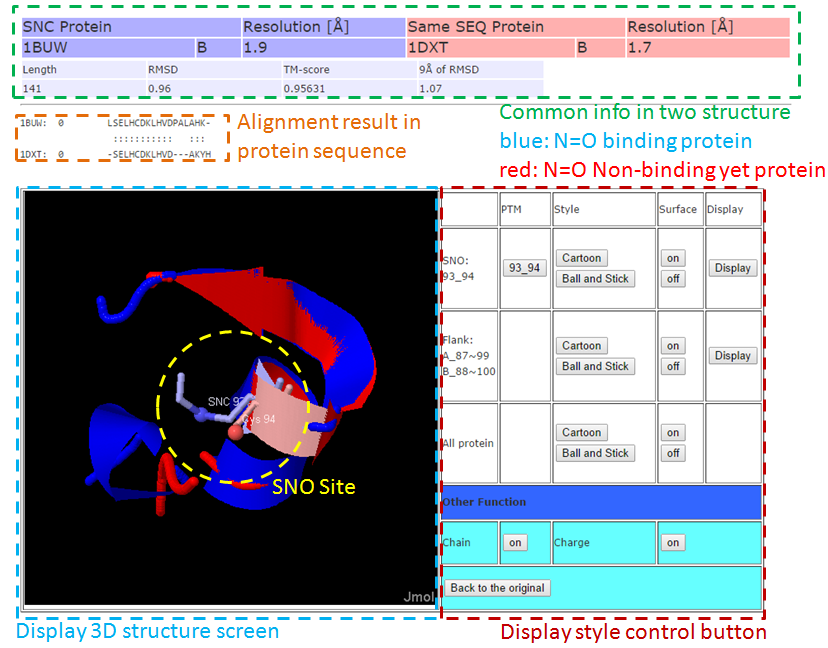
3-2. Full Structure Alignment
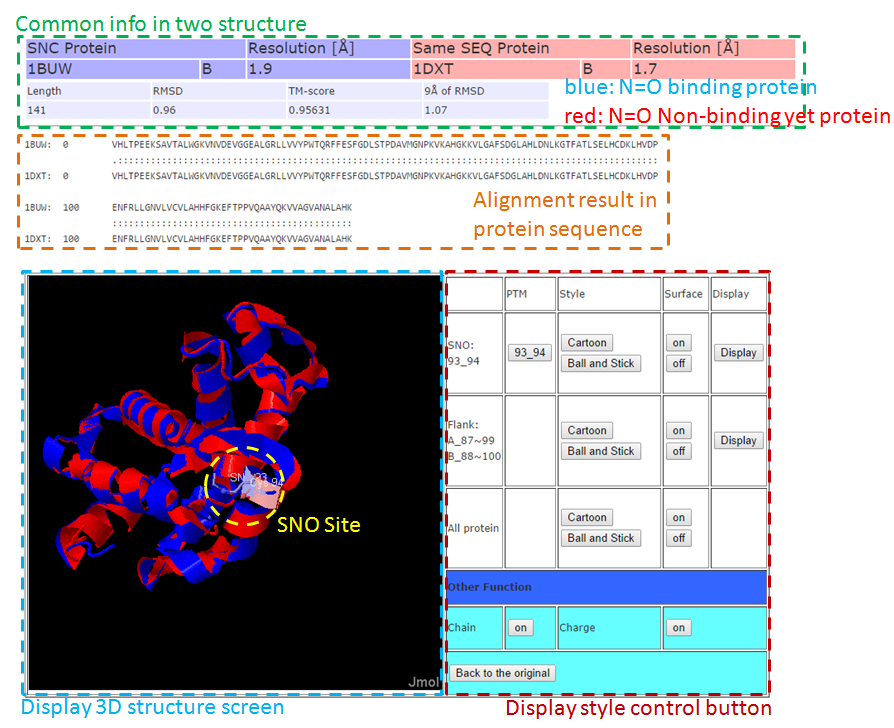
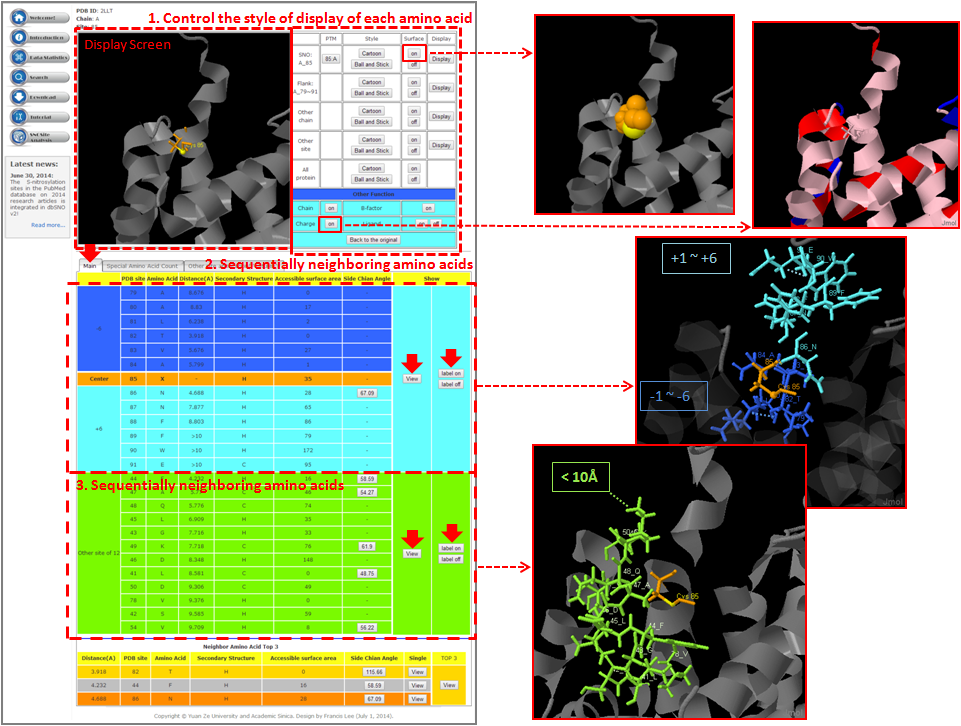
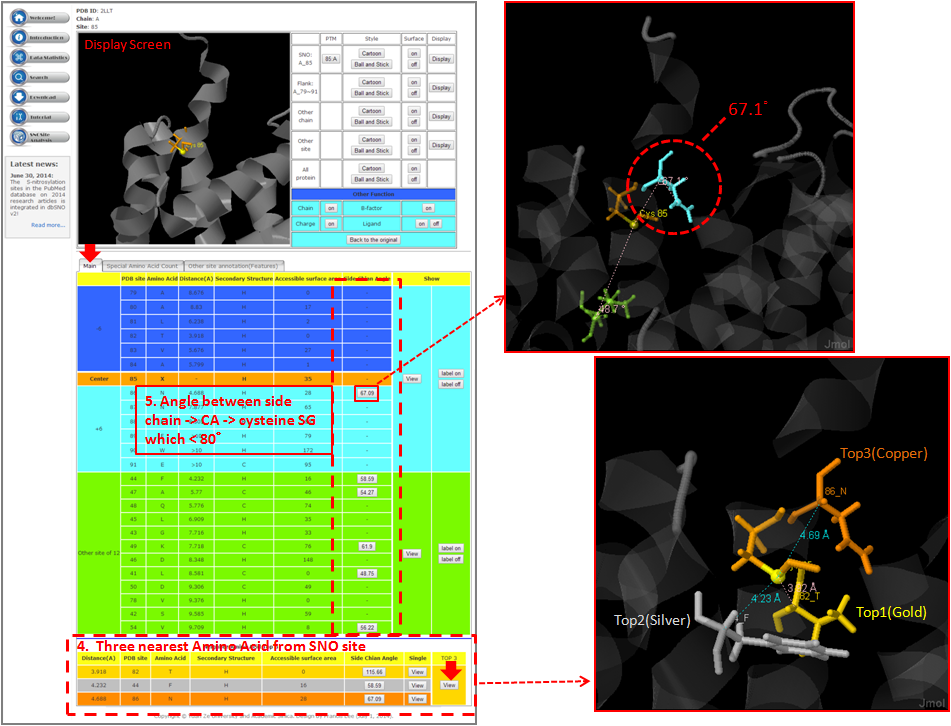
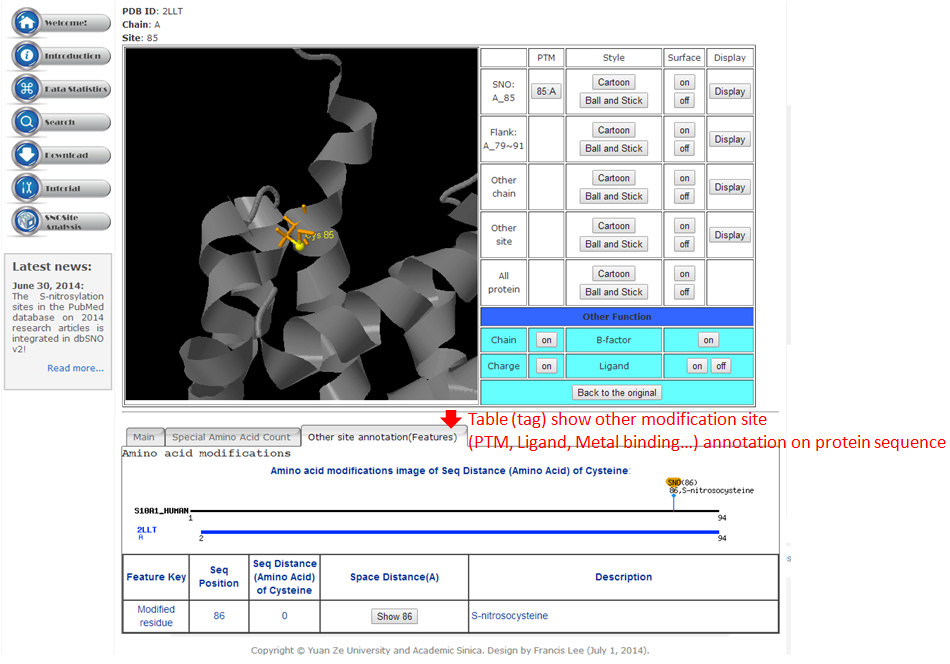
Tutorial in SNO Site Structure Analysis Page