System Flow of dbSNO 3.0
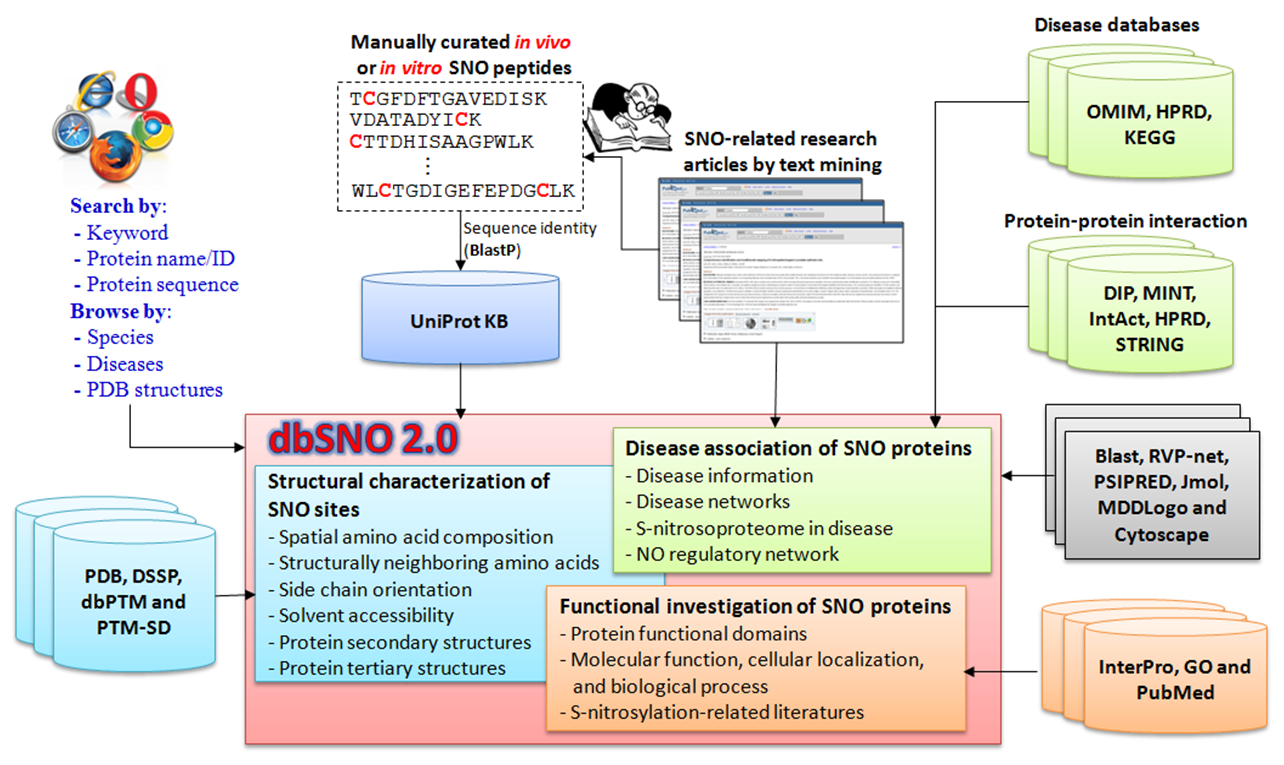
S-nitrosylation (SNO) is a reversible post-translational modification where nitric oxide (NO) is covalently attached to cysteine residues. It acts as a pleiotropic regulator, similar to phosphorylation, influencing protein function, stability, and conformation in cancers and disorders. dbSNO 3.0 is an enhanced resource for exploring SNO substrate sites and regulatory networks, mapping 4165 SNO sites across 2277 proteins to PDB entries. This version provides spatial composition, solvent-accessible surface area, and side chain orientation for 298 cysteine residues, integrating annotations for molecular functions, biological processes, and diseases. Users can search proteins/genes to reconstruct S-nitrosylation networks, with a new colorectal cancer dataset highlighting NO signaling roles. The database is freely accessible and regularly updated with new research.
| Metric | dbSNO 2.0 | dbSNO 3.0 |
|---|---|---|
| Number of organisms | 18 | 922 |
| Number of total PTM instances | 4,165 | 61,122 |
| Number of total proteins | 2,277 | 20,512 |
| Number of supported literatures | 276 | 956 |
| Number of S-nitrosylation (SNO) sites / proteins | 4,165 / 2,277 | 23,417 / 10,996 |
| Number of in vivo SNO sites / proteins | 1,169 / 768 | 1,399 / 900 |
| Number of in vitro SNO sites / proteins | 2,076 / 1,195 | 1,982 / 1,134 |
| Number of S-palmitoylation (S-Palm) sites / proteins | - | 10,239 / 5,799 |
| Number of S-glutathionylation (SSG) sites / proteins | - | 9,148 / 5,028 |
| Number of S-sulfenylation (SOH) sites / proteins | - | 4,964 / 3,280 |
| Number of S-sulfhydration (SSH) sites / proteins | - | 2,710 / 1,704 |
| Number of disulfide bond (SSR) sites / proteins | - | 9,916 / 1,544 |
| Number of S-sulfinylation (SOOH) sites / proteins | - | 728 / 524 |
| Structural characterization of cysteine modification sites | Only SNO sites | Over 6 types of cysteine PTMs |
| PTM regulatory networks | Network analysis with protein-protein interactions | Network analysis with protein-protein interactions, metabolic and signaling pathways |
| Disease annotation | KEGG Disease database, OMIM, The Human Protein Atlas | KEGG Disease database, The Human Protein Atlas, UniProt and dbPTM |
| Large language model for PTM classification and functional annotations | - | CysPTM-GPT2 predictor fine-tuned from ProtGPT2 |
| dbSNO 3.0 | All (SNO sites / proteins) | Human (SNO sites / proteins) | Mouse (SNO sites / proteins) | Rat (SNO sites / proteins) |
|---|---|---|---|---|
| In vivo | 1,399 / 900 | 958 / 594 | 425 / 293 | 0 / 0 |
| In vitro | 1,982 / 1,134 | 188 / 151 | 1,637 / 871 | 93 / 60 |
| Total | 23,417 / 10,996 | 11,946 / 4,643 | 5,478 / 2,538 | 2,290 / 1,255 |
